TCGAbiolinksGUI is stable in another field expect Transcriptome analysis. Thank you very much in advance for your time! 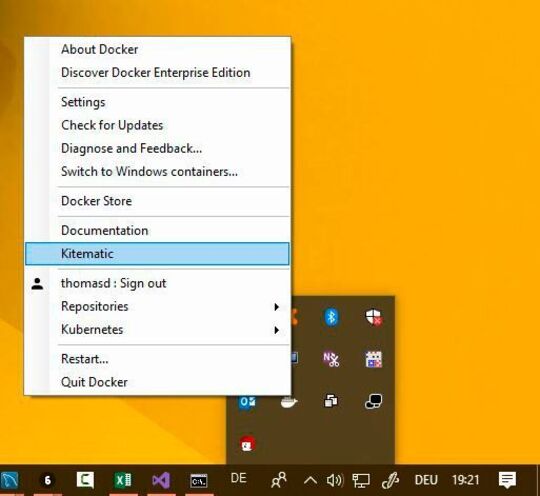 
I was wondering if my system (window7/core i3 laptop) isn't compatible with this package? I have searched and found that "stringi" is not stable in windows so I have to download and install it every session that I want to use TCGAbiolinks/GUI. Package ‘stringi’ successfully unpacked and MD5 sums checkedĬ:\Users\Sara Ansari\AppData\Local\Temp\Rtmpam8A8F\downloaded_packagesĮrror in TCGAbiolinks() : could not find function "TCGAbiolinks" Docker for windows 10 Issue 2499 docker/kitematic GitHub docker kitematic Public archive Notifications Fork Projects Closed on 41 comments Anakas commented on. There is a binary version available but the source version is later:Ĭontent type 'application/zip' length 14368013 bytes (13.7 MB) Installing package into ‘C:/Users/Sara Ansari/Documents/R/win-library/3.5’ It is a temporary solution and I have to repeat it every time I restart the Rstudio.Įrror: package or namespace load failed for ‘TCGAbiolinks’ in loadNamespace(i, c(lib.loc. I have installed it again form Github and Bioconductor websites. Warning: Error in readChar: cannot open the connectionĪnother problem is TCGAbiolinks that was working greatly is not available anymore. Suddenly, TCGAbiolinksGUI crashed and the chrome closed. It was so slow and just let me select the downloaded file. Thank you very much for your prompt reply to the questions! After installing the last package that I mentioned before, TCGAbiolinksGUI loaded perfectly and I managed to download and prepare gene expression data. I didn't change anything and immediately start working with DEA. Warning: Error in gzfile: invalid 'description' argumentħ7: deadata Ĥ9: deadata Ħ6: Warning: Error in if: argument is of length zeroĦ4: Ħ8: eval Ħ5: ĩ6: eval ĩ3: ħ5: observeEventHandler Warning: Error in if: argument is of length zeroĤ8: survivalplotdata Ĥ7: Unfortunately, I am totally new to R and do not get the warning messages! Could it be possible to interpret this message for me? I have tried to update and upgrade every package. provides: Docker Engine, Docker CLI client, Docker Compose Docker Machine, and Kitematic. Then, I tried to load it via R studio packages. In Windows 10, you install docker with Docker for Windows. After all, the chrome TCGAbiolinks was not stable and closed after a few seconds. Kitematic automates the Docker installation and setup process and provides an intuitive graphical user interface (GUI) for running Docker containers. They did not work and asked me to install stringi packages several times. Kitematic has been acquired by Docker in October 2015, so Kitematic is part of the Docker Toolbox. Kitematic is an open source project built to simplify and streamline using Docker on a Mac or Windows PC. I have tried to use docker toolbox and Kitematic software. I am trying to use TCGAbiolinksGUI on windows 7.
0 Comments
Leave a Reply. |
 RSS Feed
RSS Feed
